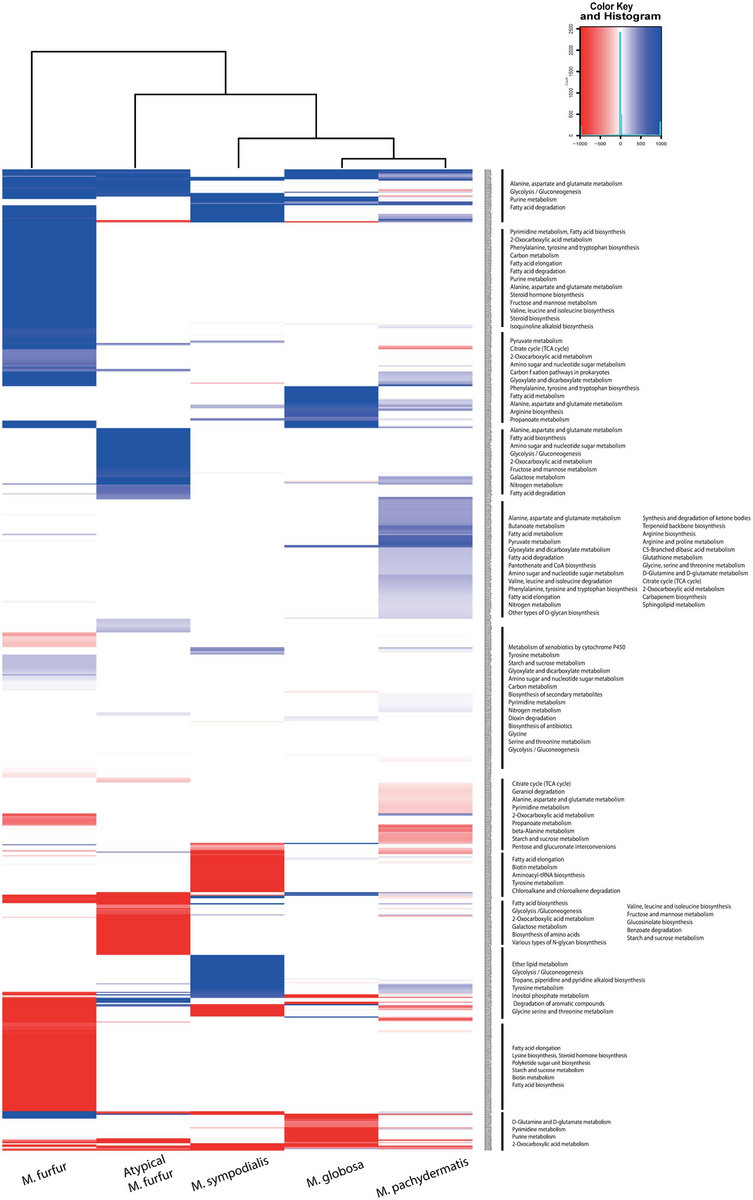

All of this known functional diversity resides in a remarkably narrow region of the family’s phylogeny ( Fig. 1) that includes all of the eukaryotic CLCs and their closest bacterial counterparts CLC-ec1, a Cl -/H + antiporter from Escherichia coli that has been subject to extensive structural and mechanistic study ( 9– 11) also resides in this region of the tree. Most CLCs thus far studied use Cl - for their physiological purposes, but has been identified as the substrate anion in a plant vacuolar CLC ( 8). CLCs participate in diverse biological tasks requiring transmembrane anion conductance, such as acidification of lysosomes, control of skeletal muscle excitability, renal regulation of blood pressure, and extreme acid resistance in enteric bacteria ( 7). This family turned out to be split into proteins of two mechanistically disparate subtypes: anion channels and Cl -/H + antiporters ( 3– 6). The CLC family of membrane proteins derives its name from its charter member, a double-barreled Cl - channel used by electric rays to generate high-power pulses to stun prey ( 1, 2). Finally, F -/H + exchange occurs with 1∶1 stoichiometry, in contrast to the usual value of 2∶1. Third, at a residue thought to distinguish CLC channels and transporters, CLC Fs bear a channel-like valine rather than a transporter-like glutamate, and yet are F -/H + antiporters. Second, CLC Fs exhibit high anion selectivity for F - over Cl. First, CLC Fs lack conserved residues that form the anion binding site in canonical CLCs. Sequence alignments and membrane transport experiments using 19F NMR, osmotic response assays, and planar lipid bilayer recordings reveal four mechanistic traits that set CLC F proteins apart from all other known CLCs. We establish here that a set of randomly selected representatives from this “CLC F” clade protect Escherichia coli from F - toxicity, and that the purified proteins catalyze transport of F - in liposomes. However these products are no longer under development.A subclass of bacterial CLC anion-transporting proteins, phylogenetically distant from long-studied CLCs, was recently shown to be specifically up-regulated by F. 
Hardware Įarly on, the company initially presented own-developed high-performance computing solutions, focusing on accelerating open source algorithms such as HMMER, Smith-Waterman and ClustalW, using FPGA technology. In 2020, CLC bio released a free plug-in that enables workflow execution on AWS directly from the CLC Genomics Workbench desktop software. In 2019, this platform was adapted for and approved for use in AWS GovCloud. 
In 2017, CLC bio launched their CLC Genomics Cloud Engine as a command-line driven platform for cloud-based bioinformatics workflow execution on Amazon Web Services. Features include read mapping and de novo assembly of high-throughput sequencing data, whole-genome detection of SNPs and structural variations, ChIP-seq, RNA-Seq, small RNA analysis, genome finishing, microbial genomics, structural biology, and functions to analyze, visualize, and compare genomic, transcriptomic, and epigenomic data. pathway analysis, genomics, and other omics). Īs additional capabilities were added to the software platform, it was eventually split into several themed Workbenches and plugins with collections of features relevant to different applications (e.g. In 2010, CLC bio was notable as the first commercial platform for bioinformatics analysis that utilized a graphical user interface for building, managing, and deploying analysis workflows as well as command-line tools, a SOAP and REST API, and later, the ability to run containerized tools. CLC bio developed some of their own open source algorithms, as well as their own SIMD-accelerated implementations of several existing popular applications. Software ĬLC bio's main activities were in software development for desktop ( Mac OS X, Windows, and Linux), enterprise, and cloud software for analysis of biological data. ĬLC bio was acquired by QIAGEN in 2013 and merged into its bioinformatics research and development division with several other purchased platforms in 2014. By 2012, it had additional offices in Cambridge, Massachusetts, Tokyo, Taipei and Delhi, with staff largely from research backgrounds (30% having a PhD) and had built a userbase of around 250,000 users in both academic institutions and biotechnology companies. Its product's development was also partly funded by collaborating with researchers on grant-funded projects. CLC bio was a bioinformatics software company that developed a software suite subsequently purchased by QIAGEN.ĬLC bio started commercial activities on Januheadquartered in Aarhus, Denmark.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
